Laboratory tests
Cross virus neutralisation (VN)
- Antigenic relatedness of different IBD viruses are calculated based on the neutralisation titres achieved with heterologous and homologous IBDV antisera.
- Using this technique Jackwood and Saif (1987) distinguished six subtypes among 13 serotype 1 IBDV strains tested.
Antigen-capture enzyme-linked immunosorbent assays (AC-ELISA)
- IBD viruses are tested in an ELISA assay against a panel of monoclonal antibodies (MCA) directed at specific defined viral epitopes.
- Viruses are then classified according to their respective MCA pattern.
- Bursal samples can be directly screened for the presence of IBDV using the AC-ELISA technique.
- Snyder et al (1988) demonstrated that classical IBD strains tested positive to MCA’s R63 and B69.
- Variant IBD strains did not test positive to MCA B69, indicating a major antigenic shift.
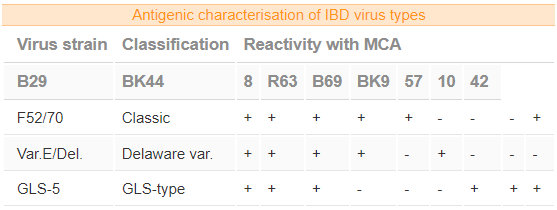
- The following table illustrates the antigenic characterisation of certain known IBD virus types against a panel of MCA’s.
RT/PCR-RFLP
(Reverse transcriptase/polymerase chain reaction-restriction fragment length polymorphism assay)
The RT/PCR-RFLP has become a popular diagnostic assay for IBDV:
- Quick detection of IBDV.
- Direct detection from bursal material.
- Detection of IBDV is possible from bursa samples preserved in a phenol and chloroform solution, allowing easy transfer of samples around the world.
The assay has two parts:
- RT/PCR: amplifying a specific fragment of the VP2 gene.
- RFLP: digestion of the amplified VP2 segment by specific restrictive enzymes. The resultant restriction fragments are then examined by gel electrophoresis.
Based on the different fragment patterns IBD viruses are identified and placed into molecular groups. Jackwood et al (2001) identified six molecular groups. IBDV strains within a group are related by ancestry.
RFLP profiles can predict the relative genetic difference between IBDV strains, however determining actual antigenic differences between viruses requires in vivo testing.
- The assay detects genetic sequences in a section of the VP2 gene that is responsible for antigenicity, thus viruses with a similar RFLP profile have the potential to be antigenically similar.
- Likewise viruses with different RFLP profiles are potentially antigenically dissimilar.
- However, some genetic changes identified by the RFLP patterns may not affect the VP2 protein sequence and thus its antigenic properties. The possibility thus exists that viruses with different RFLP profiles are antigenically similar.
- (Note: the RFLP only looks for a limited number of base pairs in the hypervariable region (HVR) of VP2 (only some cutting sites for a few enzymes which is only about 10 to 20% of the HVR). For a phylogenic tree, sequencing is required).
Peer Reviewed by Dr J J (Sjaak) de Wit and William Baxendale.

Laboratory tests for the classification of IBD virus:
- Cross virus neutralisation
- ELISA
- RT/PCR-RFLP
